Tools & Software
Open-source tools and pipelines developed throughout my research, all available on GitHub.
Developed tools
Reconstruction of synthetic tissue networks from assortativity and spatial statistics.
View on GitHub →Agent-based simulation of pancreatic ductal adenocarcinoma using the PhysiCell framework.
View on GitHub →Graphical interface for Tysserand and MOSNA — spatial transcriptomic network analysis made accessible.
View on GitHub →Nextflow pipeline for large-scale 5'UTR variant detection using MORFEE on population-scale VCF datasets.
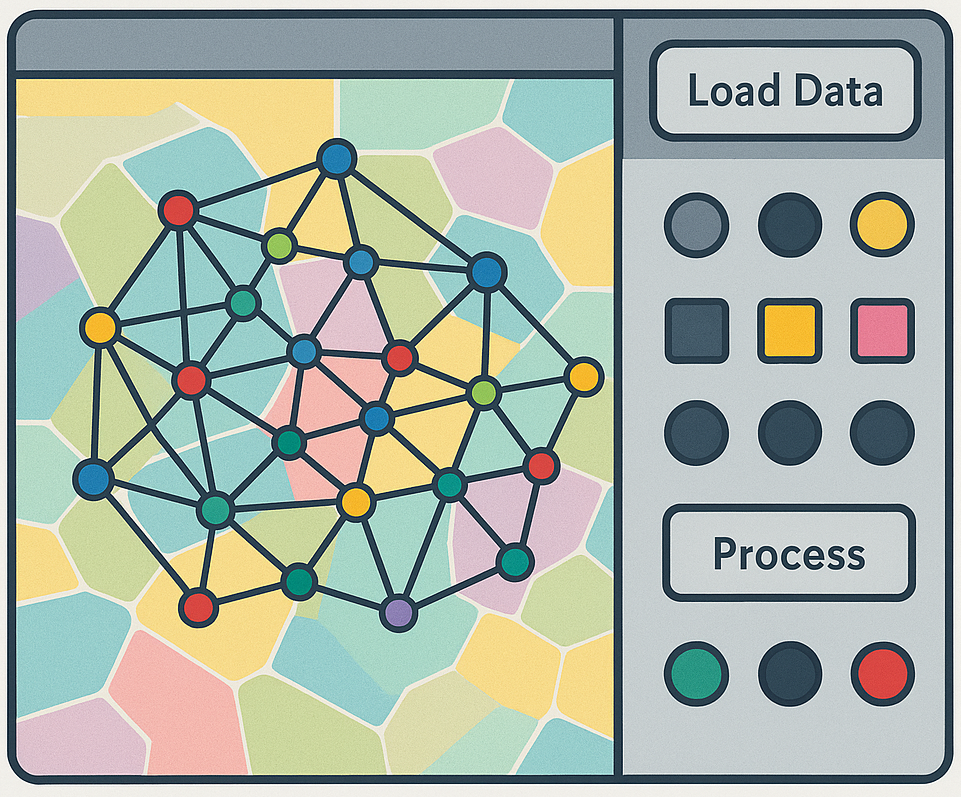
View on GitHub →MOSNA GUI
MOSNA GUI makes the MOSNA package accessible to wet-lab researchers through a user-friendly graphical interface. It wraps both Tysserand and MOSNA, allowing the configuration of parameters for spatial transcriptomic and proteomic network analysis without writing code.
The workflow covers everything from raw spatial coordinates to cell network construction, niche detection, assortativity computation, and cross-patient structural comparison — all within a single interface.
Open on GitHub